Introduction
If your application runs on Google Cloud SQL for PostgreSQL, BigQuery is usually the next step when you want fast analytics without running heavy reporting queries on the production database. In this guide, you’ll set up CDC (change data capture) so inserts, updates, and deletes in Cloud SQL PostgreSQL replicate into BigQuery with low latency.
You’ll see three practical ways teams do this:
- Google Datastream (GCP native): a serverless CDC and replication service that streams changes from operational databases into BigQuery with minimal latency.
- Estuary (managed CDC alternative): uses CDC to continuously capture updates from Cloud SQL PostgreSQL and materialize them into BigQuery tables.
- DIY Debezium (self-managed): useful if you already operate Kafka Connect and want full control (we’ll cover when it’s worth it).
What “CDC from Cloud SQL Postgres to BigQuery” actually means
PostgreSQL records changes in its WAL (write-ahead log). CDC tools read these changes using logical decoding / logical replication. In Cloud SQL, that requires enabling Cloud SQL’s logical decoding flag (cloudsql.logical_decoding) and restarting the instance so the change takes effect.
Before you start checklist (Cloud SQL, networking, BigQuery)
This checklist applies whether you use Datastream, Estuary, or a self-managed CDC pipeline. If you skip it, most failures show up later as connection errors, missing updates/deletes, or permission issues.
1) Cloud SQL PostgreSQL prerequisites for CDC
CDC tools read PostgreSQL changes from the WAL via logical decoding. In Cloud SQL, you enable this using Cloud SQL flags.
- Enable logical decoding by setting the Cloud SQL flag cloudsql.logical_decoding=on.
- Restart the Cloud SQL instance after changing the flag (the change does not take effect until restart).
- Create a dedicated user with the REPLICATION attribute for replication / CDC and follow the Cloud SQL replication setup guidance.
Practical notes that matter in real setups:
- Start with 2 to 3 tables first. Validate correctness and latency before you replicate an entire schema.
- If you expect updates and deletes to replicate correctly, make sure your tables have a primary key, or you understand how replica identity is handled (this becomes important in Datastream setups).
2) Connectivity and networking checklist
To prove the pipeline quickly, most teams start with public connectivity plus IP allowlisting and tighten security later.
- Google’s official Datastream quickstart for Cloud SQL PostgreSQL to BigQuery uses authorized networks / allowlisting as the fastest path to get the stream running.
Production hardening comes later (private networking, tighter firewalling, least privilege). The important thing is to validate CDC end-to-end first.
3) BigQuery prerequisites (dataset and permissions)
Before you turn on CDC replication:
- Create the BigQuery dataset where the replicated tables will land.
- Confirm the identity used by your chosen method can create tables (if needed) and write data into that dataset.
- This is a common setup failure point, so it’s worth checking up front.
4) Optional: If you do not need CDC (scheduled loads instead)
If you are okay with scheduled refresh (hourly or daily) rather than near real time:
- BigQuery Data Transfer Service has a PostgreSQL connector that supports PostgreSQL instances including Cloud SQL, and it runs recurring transfer jobs on a schedule.
This is useful for smaller datasets or less time-sensitive reporting, but it is not CDC.
5) Right time note
Some teams want near real-time replication 24/7. Others want to control when changes are applied in BigQuery to manage cost and warehouse load. Estuary supports configuring a sync schedule for BigQuery materializations, including sync frequency, timezone, and optional “fast sync” windows.
Method 1: (Google native) Cloud SQL PostgreSQL to BigQuery with Datastream
Datastream is Google Cloud’s serverless CDC and replication service. Google’s own docs walk through replicating Cloud SQL for PostgreSQL into BigQuery in near real time by creating a source connection profile, a BigQuery destination profile, and then a stream that ties them together.
When this method is the best fit
Choose Datastream if:
- You want a GCP-native service for CDC into BigQuery
- You prefer a managed setup (no Kafka Connect to run)
- Your primary destination is BigQuery
Google positions Datastream as a low-latency replication option into BigQuery.
Step 1: Prepare Cloud SQL PostgreSQL for CDC
Datastream reads changes using PostgreSQL logical decoding. For Cloud SQL sources, Google provides specific setup instructions.
At a minimum, you will do the following:
- Enable logical decoding for Cloud SQL PostgreSQL
- Create or configure what Datastream needs to stream changes (publication and replication slot)
- Create a user with the required permissions for CDC
Follow Google’s Cloud SQL for PostgreSQL CDC setup guide for Datastream and apply it to the database you will replicate.
Practical tip: Updates and deletes depend on the table being able to identify rows on the subscriber side. In PostgreSQL logical replication, that is based on replica identity, which is typically the primary key. If a table has no suitable replica identity, UPDATE and DELETE replication can be limited or require explicit replica identity configuration.
Step 2: Create a DataStream connection profile for the PostgreSQL source
In Datastream, create a source connection profile that points to your Cloud SQL PostgreSQL instance.
Google’s quickstart shows this flow:
- Create a PostgreSQL source connection profile
- Provide host, port, database, username, password
- Validate connectivity
Use the official quickstart steps for creating connection profiles.
Step 3: Create a Datastream connection profile for the BigQuery destination
Create a destination connection profile for BigQuery and choose how Datastream should write into BigQuery datasets.
Google documents the BigQuery destination configuration, including:
- Selecting the target BigQuery project and dataset behavior
- Write mode and staleness options
- Data type mapping considerations
Use Google’s BigQuery destination guide when selecting these settings.
Step 4: Create the stream (source to BigQuery) and select objects
Now create the stream that connects your source and destination profiles.
In the Datastream quickstart, creating a stream includes:
- Selecting the source connection profile (PostgreSQL)
- Selecting the destination connection profile (BigQuery)
- Choosing which schemas and tables to replicate
- Choosing how schemas map to BigQuery datasets (for example, one dataset per schema, or a single dataset)
Step 5: Start the stream and validate inserts, updates, deletes
Once the stream is running, validate the three operations users care about:
- Insert test
- Insert a new row into a replicated table
- Confirm the row appears in the corresponding BigQuery table
- Update test
- Update a non-key column
- Confirm the updated value shows up in BigQuery
- Delete test
- Delete the row
- Confirm it is removed (or reflected correctly, depending on the table model produced)
If you see inserts but not deletes or updates, revisit replica identity and primary key expectations. PostgreSQL logical replication requires replica identity (typically a primary key) to replicate UPDATE and DELETE operations reliably.
Operational notes that matter in production
- For PostgreSQL sources, Datastream uses a logical replication slot per stream. A large transaction or a high-churn table can delay replication for other tables in the same stream, so isolating high-volume tables into separate streams is a common scaling pattern.
- If you are using Cloud SQL for PostgreSQL as the source and BigQuery as the destination, Google also documents an “automated stream setup flow” available under specific connectivity requirements (private services access and Private Service Connect interfaces). This can reduce the number of manual steps in some environments.
Method 2: Cloud SQL PostgreSQL to BigQuery using Estuary
Estuary is the Right Time Data Platform, which means you can choose when data moves: sub second, near real time, or batch. In this method, you will:
- Capture changes from Google Cloud SQL for PostgreSQL into Estuary using CDC.
- Materialize those captured collections into Google BigQuery tables, and optionally configure a sync schedule to control how often BigQuery transactions run.
When to use this method
This approach is a good fit when you want:
- A managed CDC pipeline without stitching multiple services together
- To sync data to other destinations as well as BigQuery
- The option to control data movement timing for warehouse load or cost with a sync schedule
- To transform data before it lands in BigQuery, either through simple field selection and renaming or more advanced derived data collections
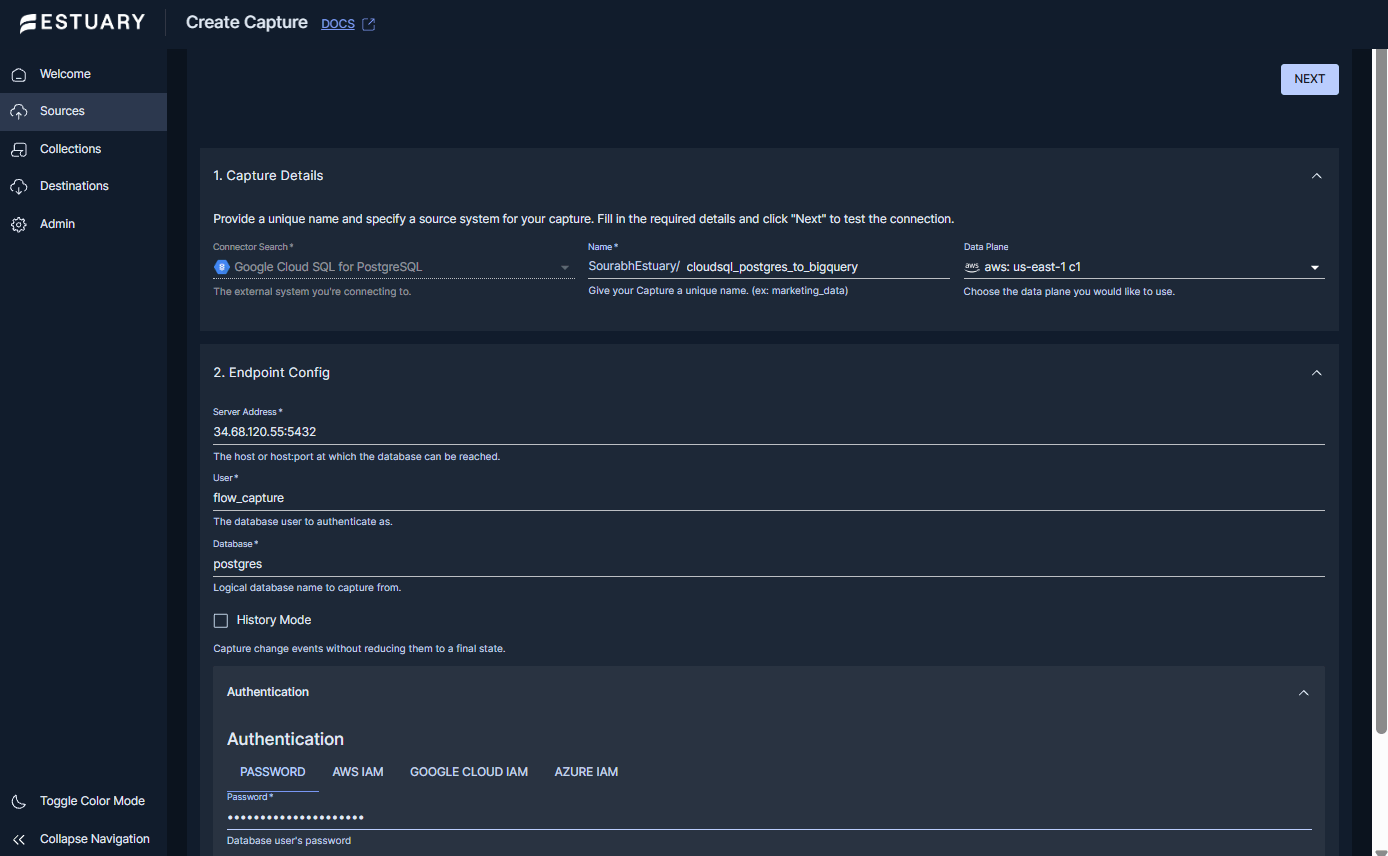
Step 1: Create a capture from Google Cloud SQL for PostgreSQL
- In the Estuary dashboard, go to Sources and select New Capture.
- Search for and select Google Cloud SQL for PostgreSQL.
- In Capture Details, provide:
- Name: a unique name for the capture.
- Data plane: choose the data plane you want to run on (select a GCP data plane in the same region as your Cloud SQL and BigQuery resources for the lowest latency and cost).
- In Endpoint Config, enter the connection details:
- Server Address: the host:port of the Cloud SQL instance.
- User: the database user to authenticate as.
- Database: the logical database name you want to capture from.
- Password or IAM Authentication: provide credentials for the Cloud SQL instance.
- (Optional) Review Advanced Options:
- Publication Name and Slot Name can be specified if you need to control which PostgreSQL publication and replication slot are used.
- SSL Mode can be set to match your Cloud SQL security requirements.
- Use Discovery Schema Selection to limit discovery to specific schemas if desired. Leaving it empty discovers tables across schemas.
- Select Read-Only Capture if your Cloud SQL instance is a read replica. Make sure you understand the caveats about this mode. Otherwise, Estuary will require write access to track progress in a watermarks table.
- Backfill-related options are available for controlling how initial history is captured.
- Click Next to test the connection and complete discovery. Then select the tables you want to capture and publish the capture.
Once published, Estuary continuously captures changes (inserts, updates, deletes) from Cloud SQL PostgreSQL into one or more collections.
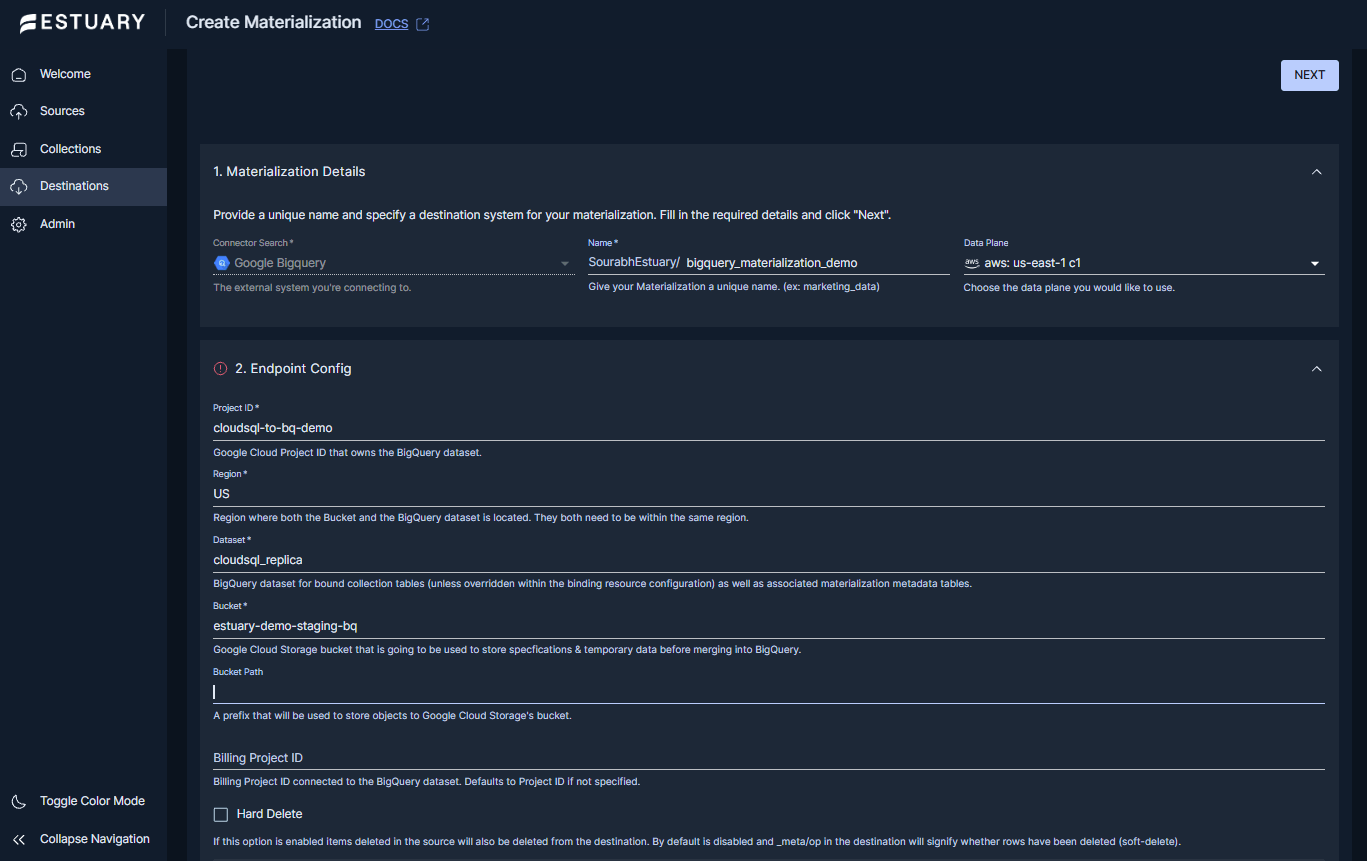
Step 2: Create a BigQuery materialization
Prerequisite: The BigQuery connector materializes via files in a Google Cloud Storage bucket (used as a temporary staging area), and the bucket must be in the same region as the destination BigQuery dataset. The service account used for authentication needs BigQuery and GCS permissions as documented in the connector prerequisites.
- Go to Destinations and select New Materialization.
- Search for and select Google BigQuery.
- In Materialization Details, provide:
- Name: a unique name for the materialization.
- Data plane: choose the data plane you want to run on.
- In Endpoint Config, configure the BigQuery destination:
- Project ID: the Google Cloud project that owns the BigQuery dataset.
- Region: the region where the dataset and bucket are located.
- Dataset: the BigQuery dataset that will contain the tables created for your collections.
- Bucket: the Google Cloud Storage bucket used by the connector.
- Authentication: provide either Service Account JSON or use GCP IAM, depending on your desired setup.
- Under Source Collections, add bindings from the captured collections to BigQuery tables. Estuary will propose bindings automatically, and you can edit naming conventions if needed.
- Publish the materialization.
At this point, Estuary is ready to write captured changes into BigQuery tables.
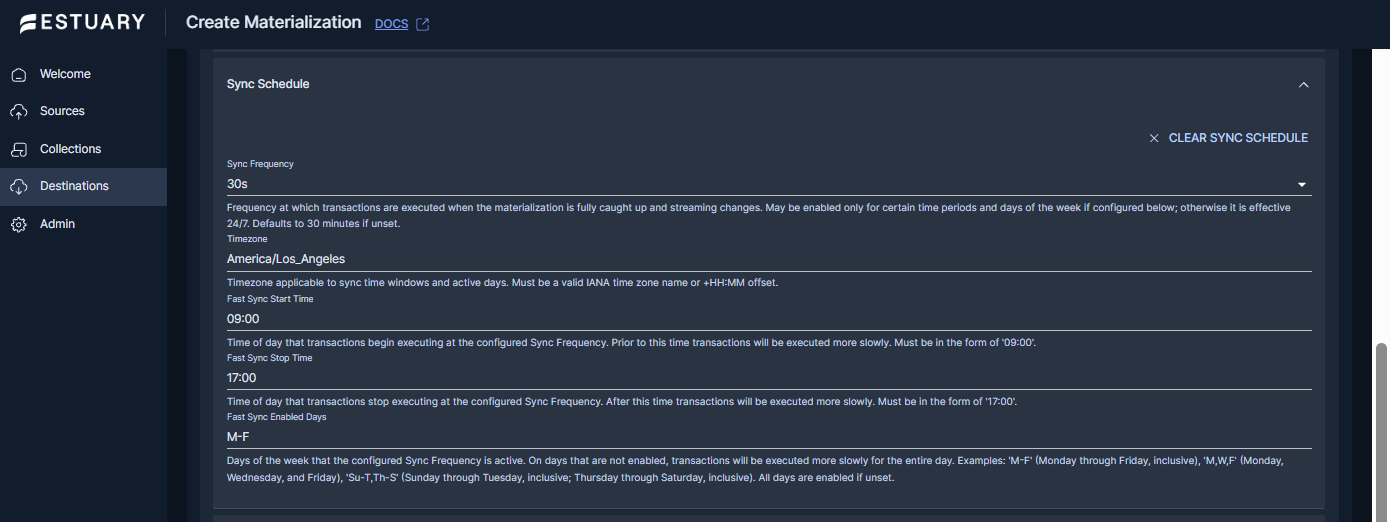
Step 3: Configure sync timing (Right Time scheduling)
For many teams, “near real time” is ideal during business hours, while off-hours updates can be slowed down to reduce warehouse load. Estuary supports a sync schedule on materializations that controls how frequently transactions execute when the materialization is caught up and streaming changes.
In the Sync Schedule section of the BigQuery materialization, configure:
- Sync Frequency: how often transactions execute.
- Timezone: the timezone used for schedule windows.
- Fast Sync Start Time and Fast Sync Stop Time: the time window where the configured sync frequency applies.
- Fast Sync Enabled Days: Which days use the fast-sync window?
Set the Sync Frequency to 0s if you want the materialization to run continuously, or set a schedule when you want predictable timing for warehouse load and cost control.
Step 4: Validate inserts, updates, and deletes in BigQuery
After publishing, validate correctness with a simple three-step test:
- Insert a row into a captured table in Cloud SQL PostgreSQL and confirm it appears in BigQuery.
- Update the row and confirm the updated value appears in BigQuery.
- Delete the row and confirm the expected delete behavior in BigQuery.
Estuary supports both soft deletes (marking documents as deleted) and hard deletes (physically removing documents), depending on how deletions are configured for the materialization.
Practical rollout tips
- Start with a small set of tables, validate correctness and latency, then expand.
- Decide whether you want “current state tables” (upserts) or a “change log” style model for auditing.
- If you expect heavy analytics use, plan your BigQuery partitioning and clustering strategy early.
Method 3: DIY pipeline with Debezium and Kafka into BigQuery
This method is for teams that want full control and already operate streaming infrastructure. The common pattern is:
Cloud SQL PostgreSQL (WAL) → Debezium PostgreSQL connector → Kafka topics → Kafka Connect sink (or custom writer) → BigQuery
Debezium turns PostgreSQL row-level changes into change events and publishes them to Kafka. For PostgreSQL, Debezium reads changes using logical decoding and a replication slot.
When DIY is worth it
Choose this approach if:
- You already run Kafka and Kafka Connect in production
- You need custom transformations in the stream before loading into BigQuery
- You are comfortable owning operational concerns like connector upgrades, schema evolution, retries, and monitoring lag
If you do not already run Kafka Connect, the operational overhead is usually higher than most teams expect.
Step 1: Configure PostgreSQL for Debezium CDC
Debezium’s PostgreSQL connector reads from the WAL using logical decoding. Depending on the output plugin you use, you may need to install a logical decoding plugin, and you will configure a replication slot and user privileges for replication.
Practical planning tips:
- Understand replication slot behavior. Slots retain WAL until consumed, which can cause WAL growth if the consumer lags.
- Start with a small set of tables and validate correctness before scaling to the full schema.
Step 2: Run Kafka Connect with the Debezium PostgreSQL connector
At a high level you will:
- Deploy Kafka and Kafka Connect
- Configure the Debezium PostgreSQL connector to publish change events into Kafka topics
- Confirm you see change events for inserts, updates, and deletes
Debezium’s official PostgreSQL connector documentation is the authoritative reference for connector properties and how logical decoding is used.
Step 3: Write changes from Kafka into BigQuery
There are two common ways to land data in BigQuery.
Option A: Use a managed BigQuery sink connector
If you are on Confluent Cloud, Confluent provides a BigQuery Sink V2 connector that uses Google’s recommended Storage Write API.
If you are running Kafka Connect yourself, Confluent also offers a self-managed BigQuery Sink connector (commonly used to stream topic data into BigQuery tables).
Important reality check for CDC:
- Debezium events are change events, not final tables. To get a clean “current state” table in BigQuery, you typically need merge logic (upserts) or a sink that applies updates and deletes correctly. This is where DIY pipelines often get complex.
Option B: Build a custom consumer using the BigQuery Storage Write API
If you need maximum control, you can consume Kafka topics and write into BigQuery using the Storage Write API. Google documents that the Storage Write API supports streaming ingestion with minimal latency, and it supports exactly-once semantics using stream offsets when you use committed streams.
This approach is powerful but you now own:
- batching, retries, idempotency
- schema mapping
- merge logic for updates and deletes (if you want current state tables)
What usually goes wrong with DIY (and why teams switch later)
These are the common failure modes that show up after the first proof of concept:
- Schema evolution overhead: coordinating schema changes across Postgres, Debezium events, converters, and BigQuery tables.
- Exactly-once expectations: BigQuery ingestion and Kafka processing semantics need careful design to avoid duplicates.
- Merge complexity: turning change events into clean current-state tables in BigQuery requires deterministic keys and consistent upsert rules.
- Lag and WAL retention risk: if CDC consumers lag, replication slots can retain WAL longer than expected.
Which method should you choose (Datastream vs Estuary vs Debezium)
All three options can replicate Cloud SQL PostgreSQL into BigQuery. The right choice usually comes down to how much infrastructure you want to operate, how flexible you need the pipeline to be, and whether you want control over timing.
Choose Datastream if you want the most GCP native path
Datastream is the clearest fit when:
- You are primarily on GCP and your destination is BigQuery
- You want a managed CDC service with minimal infrastructure
- You want a path that aligns closely with Google’s official replication guidance
Google provides an official quickstart specifically for replication from Cloud SQL PostgreSQL to BigQuery using Datastream.
Choose Estuary if you want managed CDC plus control over sync timing
Estuary is a good fit when:
- You want a managed CDC pipeline that is straightforward to set up and operate
- You want the option to control when BigQuery transactions run using a sync schedule
- You want flexibility to add more destinations later, without redesigning the pipeline
Estuary documents both the Cloud SQL PostgreSQL capture connector and the BigQuery materialization connector, and supports configuring a materialization sync schedule (frequency, timezone, and optional fast-sync windows).
Choose Debezium if you already run Kafka and need deep customization
A Debezium based pipeline is usually worth it when:
- You already operate Kafka and Kafka Connect
- You need custom event transforms before data lands in BigQuery
- You have the team bandwidth to manage connector ops, schema evolution, and merge logic
Debezium’s PostgreSQL connector documentation explains how it reads PostgreSQL changes using logical decoding and publishes change events into Kafka topics.
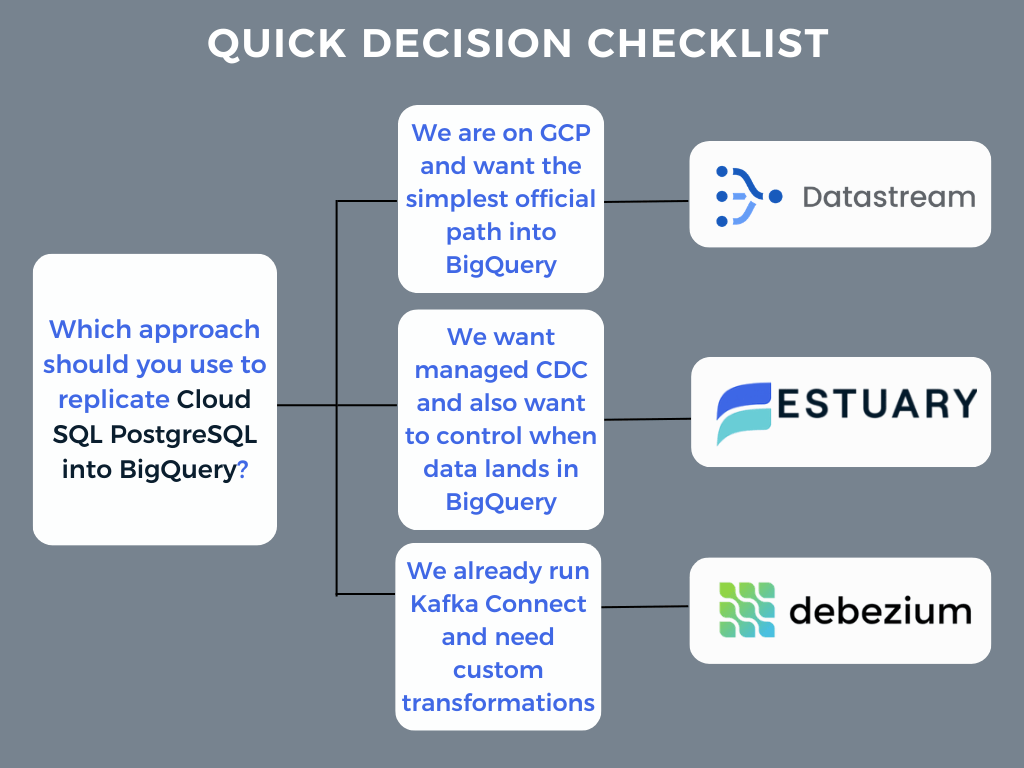
Quick decision checklist
- “We are on GCP and want the simplest official path into BigQuery” → Datastream
- “We want managed CDC and also want to control when data lands in BigQuery” → Estuary
- “We already run Kafka Connect and need custom transformations” → Debezium
BigQuery table design and performance tips for CDC workloads
CDC pipelines are only half the work. The other half is making sure the BigQuery side stays fast, affordable, and easy for analysts to use. This section covers the table patterns that work well when your source is an OLTP database (Cloud SQL Postgres) and changes are arriving continuously.
1) Pick the right table model for analytics
Most teams end up with one of these two models:
Model A: Current state tables (best for BI dashboards)
You keep one row per primary key and apply updates and deletes so BigQuery reflects the latest state. This is what most dashboard queries want.
Model B: Change log tables (best for audit and replay)
You append every change event (insert, update, delete) so you have full history. This is great for auditing, debugging, and rebuilding derived tables, but BI queries usually need an extra step to compute current state.
If your goal is self-serve analytics, current state tables are usually the default. If your goal is auditing and reproducibility, change logs are valuable.
2) Use partitioning to control cost and keep queries fast
BigQuery documentation is explicit that partitioning helps both performance and cost by reducing how many bytes a query reads when you filter on the partition column.
Practical guidance for CDC tables
- Partition tables when they are large and most queries filter by time or date.
- Choose a partition column that matches how people query, not just how data arrives.
Common partition choices:
- created_at or business event time if most analysis is time-based
- updated_at if analysts frequently look at recently changed records
- A derived date column (for example DATE(updated_at)) if that matches your BI usage
If you cannot partition on a natural event time, a safe fallback is ingestion time partitioning. The key is that dashboards should be able to prune partitions.
3) Use clustering to speed up common filters and joins
Clustered tables are sorted based on one or more columns. BigQuery notes that clustering can improve query performance and reduce cost by organizing storage blocks so BigQuery can skip more data when filters match clustered columns.
Good clustering columns for CDC workloads
Pick columns that appear constantly in filters and joins:
- customer_id, account_id, tenant_id
- user_id
- order or entity ids used in joins
A common pattern is:
- Partition by time (when possible)
- Cluster by tenant or high-cardinality business keys that your BI tool filters on
4) If you stream into BigQuery, understand the write semantics
If you are implementing your own pipeline (or evaluating one), it helps to know what BigQuery considers “at least once” vs “exactly once” for streaming ingestion.
Google’s Storage Write API docs explain:
- Use the default stream for at least once semantics (duplicates are possible)
- Use committed streams with stream offsets to achieve exactly once semantics
- Some pipelines may produce duplicates unless they apply dedupe or idempotent upserts.
- If exactness matters, confirm how the tool handles retries and offsets.
Conclusion
CDC from Cloud SQL for PostgreSQL to BigQuery is a reliable way to keep analytics fresh without running heavy queries on the production database. The key is getting the prerequisites right (logical decoding, connectivity, permissions), validating inserts, updates, and deletes early, and then tuning BigQuery tables so performance and cost stay predictable as you scale.
Next Steps and Resources
- Run the same pipeline end-to-end in Estuary
- Use the official Quickstart to create a source, capture data into collections, and materialize to a destination:
- Use the official connector docs for the exact setup in this article
- Create an account and build the pipeline on your own data
- Talk to Estuary for help with production rollout
- Browse Estuary docs by topic
FAQs
Are updates and deletes replicated, or only inserts?
Do I need a publication and replication slot for Datastream?
Can I do Cloud SQL PostgreSQL to BigQuery without CDC, on a schedule?
How do I control when data lands in BigQuery if I do not want continuous updates?

About the author
Emily is a software engineer and technical content creator with an interest in developer education. She has experience across Developer Relations roles from her FinTech background and is always learning something new.